Abstract
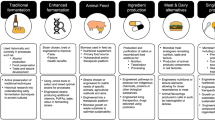
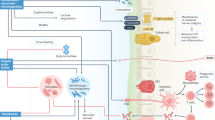
Fermented foods and beverages, produced by the intentional growth of microorganisms, have long been among the most widely consumed foods in the human diet. Whether microorganisms are added directly to food substrates, or the growth and activity of autochthonous microorganisms colonizing food substrates is encouraged by selective conditions, the production of organic acids, ethanol and other metabolic end products enhance the safety, shelf-life, sensory and functional properties of foods. Whereas some fermented foods may be produced by communities dominated by only a few taxa of limited phylogenetic diversity, others are produced through the concerted action of diverse microbial communities. In this Review, we describe the ecological interactions shaping microbial community structure and function across various categories of fermented foods by providing specific examples. We also describe how the manufacture, quality and sustainability of even traditional fermented foods can be improved by contemporary technologies. Finally, we briefly discuss current research on the ecological impact of microorganisms found in fermented food on the human gut.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
27,99 € / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
209,00 € per year
only 17,42 € per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Boethius, A. Something rotten in Scandinavia: the world’s earliest evidence of fermentation. J. Archaeol. Sci. 66, 169–180 (2016).
McGovern, P. E. et al. Fermented beverages of pre- and proto-historic China. Proc. Natl Acad. Sci. USA 101, 17593–17598 (2004).
Miller, E. R. et al. Establishment limitation constrains the abundance of lactic acid bacteria in the Napa cabbage phyllosphere. Appl. Environ. Microbiol. 85, e00269–19 (2019).
Ma, Y. et al. Microbiota dynamics and volatile metabolite generation during sausage fermentation. Food Chem. 423, 136297 (2023).
Porter, J. R. Louis Pasteur. achievements and disappointments. Bacteriol. Rev. 25, 389–403 (1961).
Marco, M. L. et al. The International Scientific Association for Probiotics and Prebiotics (ISAPP) consensus statement on fermented foods. Nat. Rev. Gastroenterol. Hepatol. 18, 196–208 (2021).
Steinkraus, K. H. in Handbook of Food and Beverage Fermentation Technology (eds Hui, Y. H. et al.) 1–8 (Marcel Dekker, 2004).
Beresford, T. P., Fitzsimons, N. A., Brennan, N. L. & Cogan, T. M. Recent advances in cheese microbiology. Int. Dairy J. 11, 259–274 (2001).
Venturini Copetti, M. Yeasts and molds in fermented food production: an ancient bioprocess. Curr. Opin. Food Sci. 25, 57–61 (2019).
Benítez-Cabello, A., Delgado, A. M. & Quintas, C. Main challenges expected from the impact of climate change on microbial biodiversity of table olives: current status and trends. Foods 12, 3712 (2023).
Choudoir, M. J., Barberán, A., Menninger, H. L., Dunn, R. R. & Fierer, N. Variation in range size and dispersal capabilities of microbial taxa. Ecology 99, 322–334 (2018).
Gomes, S. I. F. et al. Microbiota in dung and milk differ between organic and conventional dairy farms. Front. Microbiol. 11, 1746 (2020).
Streule, S., Freimüller Leischtfeld, S., Galler, M. & Miescher Schwenninger, S. Monitoring of cocoa post-harvest process practices on a small-farm level at five locations in Ecuador. Heliyon 8, e09628 (2022).
Reese, A. T., Madden, A. A., Joossens, M., Lacaze, G. & Dunn, R. R. Influences of ingredients and bakers on the bacteria and fungi in sourdough starters and bread. mSphere 5, e00950–19 (2020). Analysis of potential sources of microorganisms and their influence on composition and function of sourdough communities.
Einson, J. E. et al. A vegetable fermentation facility hosts distinct microbiomes reflecting the production environment. Appl. Environ. Microbiol. 84, e01680–18 (2018).
Bokulich, N. A., Ohta, M., Lee, M. & Mills, D. A. Indigenous bacteria and fungi drive traditional Kimoto sake fermentations. Appl. Environ. Microbiol. 80, 5522–5529 (2014).
Alexa, E. A. et al. The detailed analysis of the microbiome and resistome of artisanal blue-veined cheeses provides evidence on sources and patterns of succession linked with quality and safety traits. Microbiome 12, 78 (2024).
Hutkins, R. Microbiology and Technology of Fermented Foods 2nd edn (Wiley-Blackwell, 2018).
Wolfe, B. E. & Dutton, R. J. Fermented foods as experimentally tractable microbial ecosystems. Cell 161, 49–55 (2015).
Gänzle, M. G. et al. Starter culture development and innovation for novel fermented foods. Annu. Rev. Food Sci. Technol. 15, 211–239 (2024).
Johansen, E. Use of natural selection and evolution to develop new starter cultures for fermented foods. Annu. Rev. Food Sci. Technol. 9, 411–428 (2018).
Wolfe, B. E., Button, J. E., Santarelli, M. & Dutton, R. J. Cheese rind communities provide tractable systems for in situ and in vitro studies of microbial diversity. Cell 158, 422–433 (2014). Early example of the use of in vitro studies to gain mechanistic insights into microbial interactions important for cheese ripening.
Rousseau, G. M. & Moineau, S. Evolution of Lactococcus lactis phages within a cheese factory. Appl. Environ. Microbiol. 75, 5336–5344 (2009).
Spus, M. et al. Strain diversity and phage resistance in complex dairy starter cultures. J. Dairy Res. 98, 5173–5182 (2015). Characterization of the role of bacteriophage predation in community stability and function during the production of Gouda cheese.
Karahadian, C. & Lindsay, R. C. Integrated roles of lactate, ammonia, and calcium in texture development of mold surface-ripened cheese. J. Dairy Sci. 70, 909–918 (1987).
Galli, B. D., Martin, J. G. P., da Silva, P. P. M., Porto, E. & Spoto, M. H. F. Sensory quality of Camembert-type cheese: relationship between starter cultures and ripening molds. Int. J. Food Microbiol. 234, 71–75 (2016). Study reporting on the interplay between moulds and lactic acid bacteria and their effects on cheese properties.
Gillot, G. et al. Functional diversity within the Penicillium roqueforti species. Int. J. Food Microbiol. 241, 141–150 (2017).
Caron, T. et al. Strong effect of Penicillium roqueforti populations on volatile and metabolic compounds responsible for aromas, flavor and texture in blue cheeses. Int. J. Food Microbiol. 354, 109174 (2021).
Dalzini, E. et al. Listeria monocytogenes in Gorgonzola cheese: study of the behavior throughout the process and growth predictioni during shelf life. Int. J. Food Microbiol. 262, 71–79 (2017).
Irlinger, F., Layec, S., Hélinck, S. & Dugat-Bony, E. Cheese rind microbial communities: diversity, composition and origin. FEMS Microbiol. Lett. 362, 1–11 (2015).
Gori, K., Sørensen, L. M., Petersen, M. A., Jespersen, L. & Arneborg, N. Debaryomyces hansenii strains differ in their production of flavor compounds in a cheese-surface model. MicrobiologyOpen 1, 161–168 (2012).
Decadt, H., Vermote, L., Díaz-Muñoz, C., Weckx, S. & De Vuyst, L. Decarboxylase activity of the non-starter lactic acid bacterium Loigolactobacillus rennini gives crack defects in Gouda cheese through the production of γ-aminobutyric acid. Appl. Environ. Microbiol. 90, e0165523 (2024).
Jayabalan, R., Malbaša, R. V., Lončar, E. S., Vitas, J. S. & Sathishkumar, M. A review on kombucha tea — microbiology, composition, fermentation, beneficial effects, toxicity, and tea fungus. Compr. Rev. Food Sci. Food Saf. 13, 538–550 (2014).
Huang, X., Xin, Y. & Lu, T. A systematic, complexity-reduction approach to dissect the kombucha tea microbiome. eLife 11, e76401 (2022). Use of systems biology and synthetic communities to understand the interactions among members of the symbiotic culture of bacteria and yeasts in kombucha.
Coton, M. et al. Unraveling microbial ecology of industrial-scale kombucha fermentations by metabarcoding and culture-based methods. FEMS Microbiol. Ecol. 93, fix048 (2017).
Harrison, K. & Curtin, C. Microbial composition of SCOBY starter cultures used by commercial kombucha brewers in North America. Microorganisms 9, 1060 (2021).
Landis, E. A. et al. Microbial diversity and interaction specificity in kombucha tea fermentations. mSystems 7, e0015722 (2022).
Villarreal-Soto, S. A. et al. Metabolome-microbiome signatures in the fermented beverage, kombucha. Int. J. Food Microbiol. 333, 108778 (2020).
Blasche, S. et al. Metabolic cooperation and spatiotemporal niche partitioning in a kefir microbial community. Nat. Microbiol. 6, 196–208 (2021). Detailed analysis of microbial interactions in planktonic and sessile populations in kefir.
Fan, D., Stoyanova, L. G. & Netrusov, A. I. Microbiome and metabiotic properties of kefir grains and kefirs based on them. Microbiology 91, 339–355 (2022).
Walsh, A. M. et al. Microbial succession and flavor production in the fermented dairy beverage kefir. mSystems 1, e00052–16 (2016). Study describing the succession of microorganisms during kefir fermentation and the correlation with flavour and health.
Vegas, C. et al. Population dynamics of acetic acid bacteria during traditional wine vinegar production. Int. J. Food Microbiol. 138, 130–136 (2010).
Zhang, Z. et al. Research advances in technologies and mechanisms to regulate vinegar flavor. Food Chem. 460, 140783 (2024).
Yun, J. H., Kim, J. H. & Lee, J.-E. Surface film formation in static-fermented rice vinegar: a case study. Mycobiology 47, 250–255 (2019).
Allwood, J. G., Wakeling, L. T. & Bean, D. C. Fermentation and the microbial community of Japanese koji and miso: a review. J. Food Sci. 86, 2194–2207 (2021).
Onda, T. et al. Analysis of lactic acid bacterial flora during miso fermentation. Food Sci. Technol. Res. 9, 17–24 (2003).
Onda, T., Yanagida, F., Tsuji, M., Shinohara, T. & Yokotsuka, K. Time series analysis of aerobic bacterial flora during miso fermentation. Lett. Appl. Microbiol. 37, 162–168 (2003).
Kusumoto, K.-I. et al. Japanese traditional miso and koji making. J. Fungi 7, 579 (2021).
Koide, R. T., Kanauchi, M. & Hashimoto, Y. Variation among Japanese miso breweries in indoor microbiomes is mainly ascribed to variation in type of indoor surface. Curr. Microbiol. 81, 68 (2024).
Kothe, C. I. et al. Novel misos shape distinct microbial ecologies: opportunities for flavourful sustainable food innovation. Food Res. Int. 189, 114490 (2024). Study describing how novel miso substrates can lead to distinct microbial communities with novel properties.
Bouchez, A. & De Vuyst, L. Acetic acid bacteria in sour beer production: friend or foe? Front. Microbiol. 13, 957167 (2022).
Bongaerts, D., De Roos, J. & De Vuyst, L. Technological and environmental features determine the uniqueness of the lambic beer microbiota and production process. Appl. Environ. Microbiol. 87, e0061221 (2021). Review discussing that lambic beer fermentation relies on a succession of microorganisms that originate from the environment to produce the characteristic properties of this beer.
Spitaels, F. et al. The microbial diversity of traditional spontaneously fermented lambic beer. PLoS ONE 9, e95384 (2014).
De Roos, J. & De Vuyst, L. Microbial acidification, alcoholization, and aroma production during spontaneous lambic beer production. J. Sci. Food Agric. 99, 25–38 (2019).
Leroy, F., Verluyten, J. & De Vuyst, L. Functional meat starter cultures for improved sausage fermentation. Int. J. Food Microbiol. 106, 270–285 (2006).
Bedale, W., Sindelar, J. J. & Milkowski, A. L. Dietary nitrate and nitrite: benefits, risks, and evolving perceptions. Meat Sci. 120, 85–92 (2016).
Leroy, F., Charmpi, C. & De Vuyst, L. Meat fermentation at a crossroads: where the age-old interplay of human, animal, and microbial diversity and contemporary markets meet. FEMS Microbiol. Rev. 47, fuad016 (2023).
Liu, K. et al. Metabolic cross-feeding enhances branched-chain aldehydes production in a synthetic community of fermented sausages. Int. J. Food Microbiol. 407, 110373 (2023). Characterization of metabolic cross-feeding that occurs between bacteria and fungi during sausage fermentation.
Rong, L. et al. Fungal–bacterial mutualism: species and strain-dependent simultaneous modulation of branched-chain esters and indole derivatives in fermented sausages through metabolite cross-feeding. J. Agric. Food Chem. 72, 8749–8759 (2024).
Janßen, D., Dworschak, L., Ludwig, C., Ehrmann, M. A. & Vogel, R. F. Interspecies assertiveness of Lactobacillus curvatus and Lactobacillus sakei in sausage fermentations. Int. J. Food Microbiol. 331, 108689 (2020).
Premi, L. et al. Coagulase-negative staphylococci enhance the colour of fermented meat through a complex cross-talk between the arginase and nitric oxide synthase activities. LWT 202, 116333 (2024).
Iacumin, L. et al. Moulds and ochratoxin A on surfaces of artisanal and industrial dry sausages. Food Microbiol. 26, 65–70 (2009).
Magistà, D. et al. Penicillium salamii strain ITEM 15302: a new promising fungal starter for salami production. Int. J. Food Microbiol. 231, 33–41 (2016).
Laranjo, M., Potes, M. E. & Elias, M. Role of starter cultures on the safety of fermented meat products. Front. Microbiol. 10, 853 (2019).
Yu, A. O., Leveau, J. H. J. & Marco, M. L. Abundance, diversity and plant-specific adaptations of plant-associated lactic acid bacteria. Environ. Microbiol. Rep. 12, 16–29 (2020).
Lee, S. H., Jung, J. Y. & Jeon, C. O. Source tracking and succession of kimchi lactic acid bacteria during fermentation. J. Food Sci. 80, M1871–M1877 (2015).
Song, H. S. et al. Microbial niches in raw ingredients determine microbial community assembly during kimchi fermentation. Food Chem. 318, 126481 (2020).
Lee, S. H., Whon, T. W., Roh, S. W. & Jeon, C. O. Unraveling microbial fermentation features in kimchi: from classical to meta-omics approaches. Appl. Microbiol. Biotechnol. 104, 7731–7744 (2020).
Kim, J. Y. et al. Long-term population dynamics of viable microbes in a closed ecosystem of fermented vegetables. Food Res. Int. 154, 111044 (2022).
Galimberti, A. et al. Fermented food products in the era of globalization: tradition meets biotechnology innovations. Curr. Opin. Biotechnol. 70, 36–41 (2021).
Louw, N. L., Lele, K., Ye, R., Edwards, C. B. & Wolfe, B. E. Microbiome assembly in fermented foods. Annu. Rev. Microbiol. 77, 381–402 (2023).
Nikoloudaki, O., Aheto, F., Di Cagno, R. & Gobbetti, M. Synthetic microbial communities: a gateway to understanding resistance, resilience, and functionality in spontaneously fermented food microbiomes. Food Res. Int. 192, 114780 (2024).
Jin, R. et al. Synthetic microbial communities: novel strategies to enhance the quality of traditional fermented foods. Compr. Rev. Food Sci. Food Saf. 23, e13388 (2024).
Teusink, B. & Molenaar, D. Systems biology of lactic acid bacteria: for food and thought. Curr. Opin. Syst. Biol. 6, 7–13 (2017).
Bokulich, N. A., Thorngate, J. H., Richardson, P. M. & Mills, D. A. Microbial biogeography of wine grapes is conditioned by cultivar, vintage, and climate. Proc. Natl Acad. Sci. USA 111, E139–E148 (2014).
Ju, W. et al. Seasonal variation of metabolites in kimchi cabbage: utilizing metabolomics based machine learning for cultivation season and taste discrimination. Hortic. Environ. Biotechnol. https://doi.org/10.1007/s13580-024-00624-4 (2024).
Liu, J., Chan, S. H. J., Chen, J., Solem, C. & Jensen, P. R. Systems biology — a guide for understanding and developing improved strains of lactic acid bacteria. Front. Microbiol. 10, 876 (2019).
Hutzler, M., Morrissey, J. P., Laus, A., Meussdoerffer, F. & Zarnkow, M. A new hypothesis for the origin of the lager yeast Saccharomyces pastorianus. FEMS Yeast Res. 23, foad023 (2023).
Knudsen, S. Starters. J. Dairy Res. 2, 137–163 (1931).
Ross, R. P., Morgan, S. & Hill, C. Preservation and fermentation: past, present and future. Int. J. Food Microbiol. 79, 3–16 (2002).
Tamang, J. P. et al. Fermented foods in a global age: east meets west. Compr. Rev. Food Sci. Food Saf. 19, 184–217 (2020).
Coelho, M. C., Malcata, F. X. & Silva, C. C. G. Lactic acid bacteria in raw-milk cheeses: from starter cultures to probiotic functions. Foods 11, 2276 (2022).
Kim, D.-H., Jeong, D., Song, K.-Y. & Seo, K.-H. Comparison of traditional and backslopping methods for kefir fermentation based on physicochemical and microbiological characteristics. LWT 97, 503–507 (2018).
Leech, J. et al. Fermented-food metagenomics reveals substrate-associated differences in taxonomy and health-associated and antibiotic resistance determinants. mSystems 5, e00522–20 (2020). Analysis of metagenomes from fermented foods, sorted by substrate and composition, to identify functional traits of fermentation-associated microbiomes.
Mukherjee, A., Breselge, S., Dimidi, E., Marco, M. L. & Cotter, P. D. Fermented foods and gastrointestinal health: underlying mechanisms. Nat. Rev. Gastroenterol. Hepatol. 21, 248–266 (2024).
Valentino, V. et al. Fermented foods, their microbiome and its potential in boosting human health. Microb. Biotechnol. 17, e14428 (2024).
Hernández-Velázquez, R. et al. The future is fermented: microbial biodiversity of fermented foods is a critical resource for food innovation and human health. Trends Food Sci. Tech. 150, 104569 (2024).
Kim, E., Cho, E.-J., Yang, S.-M. & Kim, H.-Y. Identification and monitoring of Lactobacillus delbrueckii subspecies using pangenomic-based novel genetic markers. J. Microbiol. Biotechnol. 31, 280–289 (2021).
Wang, M. et al. Annotation of 2,507 Saccharomyces cerevisiae genomes. Microbiol. Spectr. 12, e0358223 (2024).
Karabín, M., Jelínek, L., Kotrba, P., Cejnar, R. & Dostálek, P. Enhancing the performance of brewing yeasts. Biotechnol. Adv. 36, 691–706 (2018).
Gibson, B. et al. Adaptive laboratory evolution of ale and lager yeasts for improved brewing efficiency and beer quality. Annu. Rev. Food Sci. Technol. 11, 23–44 (2020).
Calabrese, F. M. et al. Metabolic framework of spontaneous and synthetic sourdough metacommunities to reveal microbial players responsible for resilience and performance. Microbiome 10, 148 (2022). Comprehensive analysis of microbial interactions important for the performance of sourdough communities.
Cvetković, D. et al. An artificial neural network as a tool for kombucha fermentation improvement. Chem. Ind. Chem. Eng. Q. 28, 277–286 (2022).
Wu, L. et al. Improving the aroma profile of inoculated fermented sausages by constructing a synthetic core microbial community. J. Food Sci. 88, 4388–4402 (2023).
Jia, Y. et al. A bottom-up approach to develop a synthetic microbial community model: application for efficient reduced-salt broad bean paste fermentation. Appl. Environ. Microbiol. 86, e00306–e00320 (2020). A systems approach to improve the health benefits of a fermented food by sodium reduction, while maintaining microbiological safety.
Illikoud, N., do Carmo, F. L. R., Daniel, N., Jan, G. & Gagnaire, V. Development of innovative fermented products by exploiting the diversity of immunomodulatory properties and fermentative activity of lactic and propionic acid bacteria. Food Res. Int. 166, 112557 (2023).
Pswarayi, F. & Gänzle, M. G. Composition and origin of the fermentation microbiota of mahewu, a Zimbabwean fermented cereal beverage. Appl. Environ. Microbiol. 85, e03130–18 (2019).
Bender, D. et al. Effects of selected lactobacilli on the functional properties and stability of gluten-free sourdough bread. Eur. Food Res. Technol. 244, 1037–1046 (2018).
Vaštík, P. et al. Potential of non-Saccharomyces yeast to produce non-alcoholic beer. FEMS Yeast Res. 22, foac039 (2022).
Yabaci Karaoglan, S., Jung, R., Gauthier, M., Kinčl, T. & Dostálek, P. Maltose-negative yeast in non-alcoholic and low-alcoholic beer production. Fermentation 8, 273 (2022).
Park, J.-S. et al. Characterization of Lactobacillus fermentum PL9988 isolated from healthy elderly Korean in a longevity village. J. Microbiol. Biotechnol. 25, 1510–1518 (2015).
Pasolli, E. et al. Large-scale genome-wide analysis links lactic acid bacteria from food with the gut microbiome. Nat. Commun. 11, 2610 (2020).
David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559–563 (2014). A study providing clear evidence of the responsiveness of gut microbiota to diet, including transitory compositional changes in response to fermented foods.
Taylor, B. C. et al. Consumption of fermented foods is associated with systematic differences in the gut microbiome and metabolome. mSystems 5, e00901–e00919 (2020).
Berg, R. D. The indigenous gastrointestinal microflora. Trends Microbiol. 4, 430–435 (1996).
Savage, D. C. Microbial ecology of the gastrointestinal tract. Annu. Rev. Microbiol. 31, 107–133 (1977).
Walter, J. Ecological role of lactobacilli in the gastrointestinal tract: implications for fundamental and biomedical research. Appl. Environ. Microbiol. 74, 4985–4996 (2008).
Nielsen, E. S. et al. Lacto-fermented sauerkraut improves symptoms in IBS patients independent of product pasteurisation — a pilot study. Food Funct. 9, 5323–5335 (2018).
Cannavale, C. N. et al. Consumption of a fermented dairy beverage improves hippocampal-dependent relational memory in a randomized, controlled cross-over trial. Nutr. Neurosci. 26, 265–274 (2023).
Roselli, M. et al. Colonization ability and impact on human gut microbiota of foodborne microbes from traditional or probiotic-added fermented foods: a systematic review. Front. Nutr. 8, 689084 (2021).
Carlino, N. et al. Unexplored microbial diversity from 2,500 food metagenomes and links with the human microbiome. Cell 187, 5775–5795.e15 (2024).
Derrien, M. & van Hylckama Vlieg, J. E. T. Fate, activity, and impact of ingested bacteria within the human gut microbiota. Trends Microbiol. 23, 354–366 (2015).
Wastyk, H. C. et al. Gut-microbiota-targeted diets modulate human immune status. Cell 184, 4137–4153.e14 (2021).
Galena, A. E. et al. The effects of fermented vegetable consumption on the composition of the intestinal microbiota and levels of inflammatory markers in women: a pilot and feasibility study. PLoS ONE 17, e0275275 (2022).
Baron, M. et al. The effects of fermented vegetables on the gut microbiota for prevention of cardiovascular disease. Gut Microbiome 5, e6 (2024).
Marsh, P. D. & Percival, R. S. The oral microflora-friend or foe? Can we decide? Int. Dent. J. 56, 233–239 (2006).
Yang, I., Nell, S. & Suerbaum, S. Survival in hostile territory: the microbiota of the stomach. FEMS Microbiol. Rev. 37, 736–761 (2013).
Martinez-Guryn, K. et al. Small intestine microbiota regulate host digestive and absorptive adaptive responses to dietary lipids. Cell Host Microbe 23, 458–469.e5 (2018).
Woelfel, S., Silva, M. S. & Stecher, B. Intestinal colonization resistance in the context of environmental, host, and microbial determinants. Cell Host Microbe 32, 820–836 (2024).
McKenney, E. A. et al. Sourdough starters exhibit similar succession patterns but develop flour-specific climax communities. PeerJ 11, e16163 (2023).
Debray, R. et al. Priority effects in microbiome assembly. Nat. Rev. Microbiol. 20, 109–121 (2022).
Gänzle, M. G. & Zheng, J. Lifestyles of sourdough lactobacilli — do they matter for microbial ecology and bread quality? Int. J. Food Microbiol. 302, 15–23 (2019).
Gänzle, M. & Ripari, V. Composition and function of sourdough microbiota: from ecological theory to bread quality. Int. J. Food Microbiol. 239, 19–25 (2016).
Liu, D., Chen, Q., Zhang, P., Chen, D. & Howell, K. S. The fungal microbiome is an important component of vineyard ecosystems and correlates with regional distinctiveness of wine. mSphere 5, e00534–20 (2020).
Steenwerth, K. L. et al. Fungal and bacterial communities of ‘Pinot noir’ must: effects of vintage, growing region, climate, and basic must chemistry. PeerJ 9, e10836 (2021).
Lin, X. et al. Sodium reduction in traditional fermented foods: challenges, strategies, and perspectives. J. Agric. Food Chem. 69, 8065–8080 (2021).
Aghababaie, M., Khanahmadi, M. & Beheshti, M. Developing a kinetic model for co-culture of yogurt starter bacteria growth in pH controlled batch fermentation. J. Food Eng. 166, 72–79 (2015).
Jung, S., Hwang, I. M. & Lee, J.-H. Temperature impact on microbial and metabolic profiles in kimchi fermentation. Heliyon 10, e27174 (2024).
Pérez-Díaz, I. M. et al. in Compendium of Methods for the Microbiological Examination of Foods 4th edn (eds Downes, F. P. & Ito, K.) 521–532 (American Public Health Association, 2013).
Capece, A., Pietrafesa, R., Siesto, G. & Romano, P. Biotechnological approach based on selected Saccharomyces cerevisiae starters for reducing the use of sulfur dioxide in wine. Microorganisms 8, 738 (2020).
Shakil, M. H. et al. Nitrites in cured meats, health risk issues, alternatives to nitrites: a review. Foods 11, 3355 (2022).
Liu, M. & Nauta, A. In silico prediction of horizontal gene transfer events in Lactobacillus bulgaricus and Streptococcus thermophilus reveals protocooperation in yogurt manufacturing. Appl. Environ. Microbiol. 75, 4120–4129 (2009). A study describing how horizontal gene transfer among yoghurt microorganisms has facilitated cooperation and adaptation to the milk environment.
Kerjean, J.-R. et al. Improving the quality of European hard-cheeses by controlling of interactions between lactic acid bacteria and propionibacteria. Food Res. Int. 33, 281–287 (2000).
Sieuwerts, S., de Bok, F. A. M., Hugenholtz, J. & van Hylckama Vlieg, J. E. T. Unraveling microbial interactions in food fermentations: from classical to genomics approaches. Appl. Environ. Microbiol. 74, 4997–5007 (2008).
Landis, E. A. et al. The diversity and function of sourdough starter microbiomes. eLife 10, e61644 (2021).
Oshiro, M., Tanaka, M., Momoda, R., Zendo, T. & Nakayama, J. Mechanistic insight into yeast bloom in a lactic acid bacteria relaying-community in the start of sourdough microbiota evolution. Microbiol. Spectr. 9, e00662–21 (2021).
De Vuyst, L., Vrancken, G., Ravyts, F., Rimaux, T. & Weckx, S. Biodiversity, ecological determinants, and metabolic exploitation of sourdough microbiota. Food Microbiol. 26, 666–675 (2009).
Acknowledgements
The authors acknowledge support from the Nebraska Food for Health Center.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding authors
Ethics declarations
Competing interests
R.H. is a founder and adviser to Synbiotic Health, and J.M.A has financial interest in Synbiotic Health. H.E.H.-A. declares no competing interest.
Peer review
Peer review information
Nature Reviews Microbiology thanks Luca Coccolin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Auchtung, J.M., Hallen-Adams, H.E. & Hutkins, R. Microbial interactions and ecology in fermented food ecosystems. Nat Rev Microbiol (2025). https://doi.org/10.1038/s41579-025-01191-w
Accepted:
Published:
DOI: https://doi.org/10.1038/s41579-025-01191-w